Tonight I made a guest appearance on Steve St. Clair's SinclairDNA BlogTalkRadio episode - Pushing the Boundaries of What Can Be Learned from Your DNA.
The episode was recorded live Friday, February 21 (2014). You can listen to the recording through your computer by using this link - SinclairDNA
Steve and I talked about my recent experience with y-DNA research; Mapping y-DNA, William the Conqueror and a rare R1b group R-L11*. We'll also talked a bit on the future of y-DNA testing.
Have a listen if you get a moment. You can even download the show and take it with you.
At Origin Hunters we are passionate about DNA and helping adoptees. The answers start with your DNA.
Thursday, February 20, 2014
Monday, February 10, 2014
The Third Brother: A Y-DNA Tale
This R-L11 man had three sons, in the genetic sense, not in the literal sense. The first two sons are R-U106 and R-P312. Their stories are well known (at least in genetic genealogy circles). This is the story of the third brother, the one without a name. I’m going out on a limb in saying that this third branch exists as an independent unidentified SNP. R-DF100 has been identified as belonging to this third branch. Yet, it is too early to determine whether DF100 is the third brother or one of the many nephews (I had to keep the analogy going). Currently it is known as R-L11*/P310* (xU106,xP312), which means that folks on this branch test positive for having the L11 SNP and test negative for the U106 and P312 SNPs. Let’s call him R-x for simplicity. In case you were wondering, a SNP (single nucleotide polymorphism) is a mutation that can mark a branch point on your DNA.
 |
Figure 1 – Three Brothers
|
What do we know about R-x? They are a small group, only about 10% of the very large R1b population in Europe. They are still found in substantial numbers in Danelaw areas, the Netherlands, Pomerania, former Prussia and Denmark. U.S. President John Adams is one famous member of group R-x. A group of R-x descendants have created a site (http://www.worldfamilies.net/surnames/r1b1a2a1a) for those who are interested in tracing their family origins further back, have taken a y-DNA deep clade test and tested positive for L11 and negative for P312 / U106.
R-x was rare and it piqued my curiosity. I wanted to know how they fit into the bigger picture, where they came from and maybe connect them to a part of history. I’ve had some good success with geographical distribution of y-DNA data based on multiple distance measurements from reference positions (BGM). To start, I collected 26 R-x y-DNA records with close STR marker matches and known or probable SNP matches. Eight of these records were directly from the R1b1a2a1a website group data. The records were processed to determine time to most recent common ancestor (TMRCA). The neighbor-joining method was run on the results to create a phylogenetic tree.
 |
Figure 2 – Phylogenetic Tree
– R-L11*/P310* (xU106, xP312)
|
 |
| Figure 3 – Generalized Migration Flow – R-L11*/P310* (xU106,xP312) |
The third brother remains unnamed. Perhaps his name is R-DF100. The SNP hunters, those folks that are finding new SNPs every day, need more R-L11*/P310* (xU106,xP312) samples in order to identify a defining SNP. I’d also love to see better techniques of determining the age of a genetic branch. Someday we will know the name and the birthdate of the third brother.
Reference:
Maglio, MR (2014) Y-Chromosome Haplotype Origins via Biogeographical Multilateration (Link)
© MRMaglio 2014
Maglio, MR (2014) Y-Chromosome Haplotype Origins via Biogeographical Multilateration (Link)
© MRMaglio 2014
Wednesday, January 29, 2014
Getting More from Your Genetic Testing Results: Y-DNA Mapping
Y-DNA testing has only been around for about 15 years and can tell
us about our direct paternal ancestry.
It is sometimes criticized for only being able to tell us about a small
portion of our genetic history. I think
the critics forget that our mothers have fathers and there are a number of ways
to obtain male test subjects to expand Y results across the entire family
tree. In the last few years, there has
been a brighter spotlight on autosomal testing and its ability to test a larger
portion of our genes. This may have
diverted research attention. Relatively
speaking, y-DNA is still in its infancy and there is much more to be learned. In addition to our deep ancestral origins
dating back over 10,000 years, we should be able to identify our old world
homeland and our nomadic ancestor’s migration routes. It is meaningful to know not only where they
lived, but also when they lived to give us clues to their part in history.
My closest genetic cousins and I share a common
ancestor over 1,100 years ago. With that many years between us, I didn’t
spend a lot of time looking for a common surname. For the last four
years, I’ve been working with y-DNA to determine what can be learned beyond
cousin matches and distant origins. I originally tested with the
Genographic Project and then transferred my results to Family Tree DNA for an
upgrade. In the time since getting my upgrade to 67 markers, I have
received only one match. This is due to my haplotype being somewhat rare
and not a reflection of Family Tree DNA’s ability to provide matches. I learned
that I was part of haplogroup G, from the Caucasus Mountains. I
didn’t learn anything about how the ancestors of my Italian family traveled
from Western Asia to Italy.
I needed to find more cousin matches than the client
base that had tested with FTDNA. Ancestry.com allowed me to enter my
y-DNA markers into their DNA section. I compared my haplotype against the
database at Sorenson Molecular (SMGF.org). I also tried Genebase.com and
any other place I could find. I didn’t locate any cousins. I did
get a good education on what is out there for DNA companies and services.
FTDNA allowed me to transfer my results to Ysearch.org, a free public DNA
matching service that they provide. Ysearch provided much more
flexibility. Where FTDNA would only show me matches with a genetic
distance of seven or less, Ysearch allowed me to select the genetic distance.
Without the genetic distance restrictions, Ysearch showed me hundreds of very
distantly related cousins. I’d rather have distant than none. FTDNA
is not trying to be difficult. They are trying to be realistic and keep
matches within genealogical timeframes. I just needed more.
While not used consistently, one of the best features
of Ysearch is its ability to present the most distant known paternal ancestor
(MDKPA) and their origin. So now, I had cousins and their ancestral
origins. The greater the genetic distance, the closer I got to the
Caucasus Mountains. I mapped my closest cousins and their pushpins
created a line from the Alps to the North Sea, directly along the Rhine River.
This was a pattern. I like patterns. There is usually a
scientific reason for a pattern.
Figure 1 – Mapping Ancestral Origins
I’ve run a few hundred
genetic mapping exercises on the majority of Y haplogroups. The patterns are consistent. Our ancestors traveled along rivers and
coastlines. They skirted mountain ranges
and crossed bodies of water. There is
always a flow to the patterns, but the science and the why were still missing. I’ve always been interested in early Eurasian
history and have tried to associate it to the patterns in the maps that I
generated. Some maps have places and
dates that connect well to notable events.
Other maps raise more questions than answers. Genetic tests and history alone were not
making a complete picture. I needed to
add population genetics, migration models, anthropology and evolutionary
biology to my knowledge base.
Take any two people on the planet, compare their DNA and you can
calculate, approximately, how far back in time their common ancestor lived –
time to most recent common ancestor (TMRCA).
Our ancestors were nomadic and traveled about 25 to 30 km per generation
or roughly 1 km/year on average. If a
common ancestor lived 300 years ago, then that person’s descendants may have
migrated 300 km from the geographic origin of that ancestor. In the figure below, d1 and d2
represent the origins of two known y-DNA genetic records. The circles show the distance their ancestors
may have migrated. The intersections are
the potential locations of their common ancestor. In this case, there are two intersections, a1
and a2. More y-DNA records
are required to figure out which intersection is correct.
Figure 2 – Distance to Most Recent Common Ancestor
(Bilateration)
The genetic analysis that
I have developed is similar to a navigation technique. Radio navigation uses two or more beacons
with known locations and a measurement of the time it takes to receive a signal
from each. The time is converted to a
distance. A current location can then be
identified. In my method, the “beacons”
are the ancestral geographic origins. The
“signal” is the TMRCA, measured in years, converted to a distance by
multiplying the average migration rate – creating a distance to most recent
common ancestor (DMRCA). A location for
the common ancestor can then be figured out by looking at the intersections. Multiple y-DNA records are needed to
determine migration direction and geographic origins of a related set of
genetic cousins. The science is starting
to take shape.
I am G-Z726, which is a subgroup of G-Z725. At Ysearch or on an FTDNA project you may still see an older naming convention - G2a3b. As an example, let’s look
at 18 of my closest genetic cousins – haplogroup G-Z725, (DYS388=13). TMRCA data is generated using Dean McGee’s
Y-Utility. The output is turned into a
phylogenetic tree and the DMRCA is calculated.
Figure 3 - Phylogenetic Tree - G-Z725 Sample
Each distance is used to draw a circle that represents the
migration range of that ancestral line.
My range (MAG) and the range of my next closest cousin on the tree (BAB)
create two intersections on the map. One
point is in Africa and
Figure 4 - Intersection of MAG and BAB
By adding the range of cousin EBE, the correct intersection representing
common ancestor 7 (CA7) is identified.
Pairs of records are mapped to continue the analysis.
Figure 5 - Intersection of BIR and RUF
BIR and RUF are another example and they identify common ancestor
CA3. Each record is added until all
common ancestors are identified.
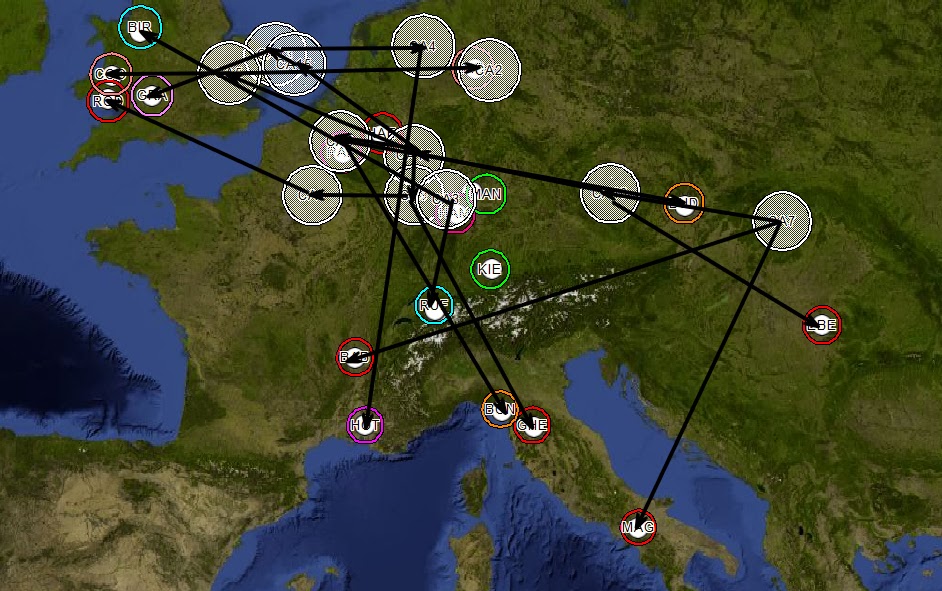
Figure 6 - Common Ancestors Mapped
Connections are drawn based on the phylogenetic tree.
Figure 7 - Phylogenetic Tree Superimposed
Then for simplification, the migration flow is generalized. Based on the phylogenetic analysis, common
ancestor 7 (CA7) is the most distant common ancestor (MDCA) of samples in this
example. The western migration flow has
a slight correlation to the Danube River and the North/South migrations have a
strong correlation to the Rhine River.
Early attempts at direct mapping of genetic data (Fig. 1) gave similar
results based on TMRCA alone. There was
no corroborating
Figure 8 - Generalized Migration Flow - G-Z725
The new methodology that I have developed
shows ancestral origins and migration direction. I call this method biogeographical
multilateration (BGM). It is the
geographical distribution of y-DNA data based on multiple distance measurements
from reference positions.
This method also has the
potential to identify the location of a DNA sample when the origin is unknown. For
one of my clients, I identified that his paternal ancestor that had immigrated
to the United States may have anglicized their name. The previous name was German and that surname
was found predominantly in central Germany.
Applying biogeographical multilateration to my client’s y-DNA results,
while leaving the origin as an unknown, indicated a continental European origin
within 200km of the city with the highest surname density.
In the referenced paper,
the example haplogroup, I-L22, has an approximate correlation to the
Norse/Viking invasions of Britain and identifies a Frisian coast staging area.
Figure 9 - Generalized Migration Flow - I-L22
We are still in the
infancy of what we can learn from y-DNA.
My BGM analysis has room for refinement.
There is enough science out there to help us. New research is being done at the aggregate
population level. We can apply elements
of that research to the individual level.
Biogeographical multilateration has the potential to fill the gap in our
knowledge between genealogical records and deep haplogroup roots. This new tool can give us old world ancestral
origins, migration flow and historical timeframes.
I needed to learn more
about my DNA and my origins. The big
testing companies haven’t filled the knowledge gaps that exist. They are in the business of providing quality
test results. The analysis and tool
creation is falling on the shoulders of “citizen scientists”. Sometimes, you just have to do it yourself.
Reference:
Maglio, MR (2014) Y-Chromosome
Haplotype Origins via Biogeographical Multilateration (Link)
Sunday, January 19, 2014
Pandora's DNA
Deep Into DNA*
Ah, poor Pandora. So slandered.
Ah, poor Pandora. So slandered.
As the story goes, Zeus gave her a jar and told her never to open it. Pandora’s curiosity got the better of her and she released all the ‘evils’ upon mankind. This is one of many origin stories for why there is evil in the world. This is also a metaphor for the spread or release of information. Too much knowledge can be ‘evil’.
New knowledge discoveries are often also slandered. DNA test results are a modern example of disrupting the status quo.
...continued at The In-Depth Genealogist with a free membership.
*The Deep Into DNA article series is published each month in the new Going In-Depth
digital genealogy magazine presented by The In-Depth Genealogist.
#gDNA
*The Deep Into DNA article series is published each month in the new Going In-Depth
digital genealogy magazine presented by The In-Depth Genealogist.
#gDNA
Sunday, July 21, 2013
Conquering William's DNA
One of my favorite aspects of y-DNA is that it’s used to prove or disprove that two men with the same last name are closely related. Two family lines with a similar surname can figure out if they have a common ancestor. The DNA matches or it doesn’t. What do you do if the common ancestor you are looking for doesn’t have a surname? If you are researching the British Isles, the surname you are looking for is probably less than 1,000 years old.
What were the surnames associated with William the Conqueror? To start, who was William the Conqueror? William the ‘bastard’ was born about 1028 in Normandy, the illegitimate son of Robert I, Duke of Normandy, and Herleva. William was the 3rd great grandson of Rollo, the Viking who harassed the French so much that they gave him Normandy in order to make him stop.
In 1066, when King Edward ‘the confessor’ of England died, William was a potential heir to the English crown. When he didn’t get the nod, he took the crown by force by defeating and killing King Harold at the Battle of Hastings.
Finding the DNA of William the Conqueror is not that easy. He has no documented living male-line descendants. King Henry I was his last legitimate offspring. If you look in the phone book, you won’t find too many names listed under Conqueror, William T. That makes asking for a DNA sample problematic.
We have to look at the entire line of Dukes from the House of Normandy to identify the surnames that they would eventually adopt. The line from Rollo to William looks like this – Rollo (846-931) > William I (900-942) > Richard I (933-996) > Richard II (978-1026) > Robert I (1000-1035) > William II (1028-1087). To start, there is some evidence, true or not, that the surnames Clifford, Devereaux and St. Clair have a direct connection back to Richard I and Richard II. It’s not my goal to prove anyone’s genealogy. Many medieval genealogies are pure fiction, geneamyth. Although, with ever story there may be a piece of the truth. Some of William’s companions at the Battle of Hastings were his cousins and it would have made sense for him to surround himself with kin. I collected those names and others that had a tenuous connection.
I began the process with the following 27 names; Bartelott, Beaumont, Bruce, Clifford, Corbett, D’Arcy, Devereaux, Giffard, Hereford, Lindsay, Molyneaux, Montgomery, Mortimer, Mowbray, Neville, Norman, Norton, Osbern, Pearsall, Ramsey, Spencer, St. Clair, Stewart, Sutton, Talbott, Umfreville and Warren. While this is not an exhaustive list, it did provide 3,800 records to sift through.
DNA records for these surnames were collected from publically available sources and sorted into haplogroups. Remember, everyone is related. It’s just a question of how far back in time they share a common ancestor. Members of haplogroups I and J may share an ancestor about 30,000 years ago, but my goal is to find as many surnames that have a common ancestor about 1,000 years ago. So, DNA comparison was limited to within haplogroups. Immediately, groups E1b, G2a, I2, J and R1a were eliminated for having no cross surname relationships.
The first likely candidate was haplogroup I1. I1 would make sense. It is a typical Scandinavian group and Rollo is supposed to be either Norwegian or Danish. There was some good cross surname relationships among 8 of the 27 surnames. More analysis showed that they didn’t form a tight clan and that their common ancestor would have been over 1,250 years ago. That predates Rollo. This doesn’t completely rule out haplogroup I1, but my expectation was that there would be a higher number of surnames and a common ancestor between Rollo and William.
The next candidate was group R1b, the most populous haplogroup in Europe and having a potential Scandinavian or continental Europe origin. This group clustered well across 25 of the 27 surnames and revealed a genetically related clan. To make sure that this wasn’t a false positive or something symptomatic about the large R1b population, I took a random sample of British Isles R1b y-DNA and ran the same comparison. The random sample did not group well and actually formed multiple clusters.
This looks very positive for the R1b group. Twenty-one of the surnames are tightly related enough that their common ancestor lived 1,080 years ago (933 AD), coincidentally the birth year of Richard I. All common ancestor calculations come with a margin of error. I’d say this estimate is plus or minus a generation. Clifford, Devereaux and St. Clair, with their genealogical connection remain in this group as well as Beaumont, Giffard, Montgomery, Mortimer, Osbern and Warren.
The odd thing about this second group of names is that they all, genealogically, connect back to Gunnora, wife/concubine of Richard I. Beaumont and Giffard are descendants of Duvelina, a sister of Gunnora. Osbern is a descendant of Herfast, a brother of Gunnora. None of this common y-DNA came from Gunnora or her sister; being female, they don’t have y-DNA to pass down. We have to look for a common male donor. My theory is that the practice of droit du seigneur – ‘right of the lord’ or primae noctis – ‘right of the first night’ was being used by Richard to increase his genetic success.
Do you have a connection to William the Conqueror? There is an estimate that 25% of the population of England is related to Bill the Conq. From a y-DNA perspective, this percentage would be lower. If you have one of these surnames; Bartelott, Beaumont, Bruce, Clifford, Corbett, D’Arcy, Devereaux, Giffard, Molyneaux, Montgomery, Mortimer, Norton, Osbern, Pearsall, Ramsey, Spencer, St. Clair, Stewart, Talbott, Umfreville (Humphrey) or Warren and match the 37-marker William the Conqueror Modal Haplotype (WCMH), you may be related.
You might match the WCMH within a few steps and not have one of those surnames. The wealthy practiced polygyny. They had as many mistresses as they could afford. The illegitimate male offspring would have generated countless undocumented surnames and carry these same y-DNA markers.
I can’t say that this is exactly William the Conqueror’s y-DNA markers. These values are a mode, the numbers that appear most frequently in the related R1b sample of 152 records. The results that I have found are based on my analysis of about 3,800 y-DNA samples and form a good correlation. New data in the future may change the results.
The techniques that I have used are similar to the ones used to identify Carthaigh (McCarthy King of Desmond), Niall of the Nine Hostages and Genghis Khan. I predict that as the DNA databases grow, more discoveries like this will be found. My next projects are to determine Rollo’s origin (see Exploring Rollo...) and Charlemagne’s haplotype.
Reference:
Maglio, MR (2013) A Y-Chromosome Signature of Polygyny in Norman England (
Link)
©Michael R. Maglio and OriginsDNA
What were the surnames associated with William the Conqueror? To start, who was William the Conqueror? William the ‘bastard’ was born about 1028 in Normandy, the illegitimate son of Robert I, Duke of Normandy, and Herleva. William was the 3rd great grandson of Rollo, the Viking who harassed the French so much that they gave him Normandy in order to make him stop.
In 1066, when King Edward ‘the confessor’ of England died, William was a potential heir to the English crown. When he didn’t get the nod, he took the crown by force by defeating and killing King Harold at the Battle of Hastings.
Finding the DNA of William the Conqueror is not that easy. He has no documented living male-line descendants. King Henry I was his last legitimate offspring. If you look in the phone book, you won’t find too many names listed under Conqueror, William T. That makes asking for a DNA sample problematic.
We have to look at the entire line of Dukes from the House of Normandy to identify the surnames that they would eventually adopt. The line from Rollo to William looks like this – Rollo (846-931) > William I (900-942) > Richard I (933-996) > Richard II (978-1026) > Robert I (1000-1035) > William II (1028-1087). To start, there is some evidence, true or not, that the surnames Clifford, Devereaux and St. Clair have a direct connection back to Richard I and Richard II. It’s not my goal to prove anyone’s genealogy. Many medieval genealogies are pure fiction, geneamyth. Although, with ever story there may be a piece of the truth. Some of William’s companions at the Battle of Hastings were his cousins and it would have made sense for him to surround himself with kin. I collected those names and others that had a tenuous connection.
I began the process with the following 27 names; Bartelott, Beaumont, Bruce, Clifford, Corbett, D’Arcy, Devereaux, Giffard, Hereford, Lindsay, Molyneaux, Montgomery, Mortimer, Mowbray, Neville, Norman, Norton, Osbern, Pearsall, Ramsey, Spencer, St. Clair, Stewart, Sutton, Talbott, Umfreville and Warren. While this is not an exhaustive list, it did provide 3,800 records to sift through.
DNA records for these surnames were collected from publically available sources and sorted into haplogroups. Remember, everyone is related. It’s just a question of how far back in time they share a common ancestor. Members of haplogroups I and J may share an ancestor about 30,000 years ago, but my goal is to find as many surnames that have a common ancestor about 1,000 years ago. So, DNA comparison was limited to within haplogroups. Immediately, groups E1b, G2a, I2, J and R1a were eliminated for having no cross surname relationships.
The first likely candidate was haplogroup I1. I1 would make sense. It is a typical Scandinavian group and Rollo is supposed to be either Norwegian or Danish. There was some good cross surname relationships among 8 of the 27 surnames. More analysis showed that they didn’t form a tight clan and that their common ancestor would have been over 1,250 years ago. That predates Rollo. This doesn’t completely rule out haplogroup I1, but my expectation was that there would be a higher number of surnames and a common ancestor between Rollo and William.
The next candidate was group R1b, the most populous haplogroup in Europe and having a potential Scandinavian or continental Europe origin. This group clustered well across 25 of the 27 surnames and revealed a genetically related clan. To make sure that this wasn’t a false positive or something symptomatic about the large R1b population, I took a random sample of British Isles R1b y-DNA and ran the same comparison. The random sample did not group well and actually formed multiple clusters.
The odd thing about this second group of names is that they all, genealogically, connect back to Gunnora, wife/concubine of Richard I. Beaumont and Giffard are descendants of Duvelina, a sister of Gunnora. Osbern is a descendant of Herfast, a brother of Gunnora. None of this common y-DNA came from Gunnora or her sister; being female, they don’t have y-DNA to pass down. We have to look for a common male donor. My theory is that the practice of droit du seigneur – ‘right of the lord’ or primae noctis – ‘right of the first night’ was being used by Richard to increase his genetic success.
Do you have a connection to William the Conqueror? There is an estimate that 25% of the population of England is related to Bill the Conq. From a y-DNA perspective, this percentage would be lower. If you have one of these surnames; Bartelott, Beaumont, Bruce, Clifford, Corbett, D’Arcy, Devereaux, Giffard, Molyneaux, Montgomery, Mortimer, Norton, Osbern, Pearsall, Ramsey, Spencer, St. Clair, Stewart, Talbott, Umfreville (Humphrey) or Warren and match the 37-marker William the Conqueror Modal Haplotype (WCMH), you may be related.
|
DYS393
|
DYS390
|
DYS19
|
DYS391
|
DYS385a
|
DYS385b
|
DYS426
|
DYS388
|
DYS439
|
DYS389i
|
DYS392
|
DYS389ii
|
|
13
|
24
|
14
|
11
|
11
|
14
|
12
|
12
|
12
|
13
|
13
|
29
|
|
DYS458
|
DYS459a
|
DYS459b
|
DYS455
|
DYS454
|
DYS447
|
DYS437
|
DYS448
|
DYS449
|
DYS464a
|
DYS464b
|
DYS464c
|
DYS464d
|
|
17
|
9
|
10
|
11
|
11
|
25
|
15
|
19
|
29
|
15
|
15
|
17
|
17
|
|
DYS460
|
Y-GATA-H4
|
YCAIIa
|
YCAIIb
|
DYS456
|
DYS607
|
DYS576
|
DYS570
|
CDYa
|
CDYb
|
DYS442
|
DYS438
|
|
11
|
11
|
19
|
23
|
15
|
15
|
17
|
17
|
36
|
37
|
12
|
12
|
You might match the WCMH within a few steps and not have one of those surnames. The wealthy practiced polygyny. They had as many mistresses as they could afford. The illegitimate male offspring would have generated countless undocumented surnames and carry these same y-DNA markers.
I can’t say that this is exactly William the Conqueror’s y-DNA markers. These values are a mode, the numbers that appear most frequently in the related R1b sample of 152 records. The results that I have found are based on my analysis of about 3,800 y-DNA samples and form a good correlation. New data in the future may change the results.
The techniques that I have used are similar to the ones used to identify Carthaigh (McCarthy King of Desmond), Niall of the Nine Hostages and Genghis Khan. I predict that as the DNA databases grow, more discoveries like this will be found. My next projects are to determine Rollo’s origin (see Exploring Rollo...) and Charlemagne’s haplotype.
Reference:
Maglio, MR (2013) A Y-Chromosome Signature of Polygyny in Norman England (
Link)
©Michael R. Maglio and OriginsDNA
Monday, March 25, 2013
The DNA of Thomas Jefferson: [Insert Shocking Title Here]
Thomas Jefferson was part of the rare y-DNA haplogroup T (formerly K2) and much has been written about his genetics. Not everything written has been logical in its assumptions. Sometimes the story lines misinterpret the underlying science.
“Was Thomas Jefferson the first Jewish President?”
“Thomas Jefferson was Phoenician.”
“If Jefferson was Phoenician, then Charlemagne was also.”
“Thomas Jefferson could have recent origins in the Middle East.”
“Thomas Jefferson’s DNA traced back to Egypt.”
In these situations the writers took a single data point and ran with it out of context.
When we look at Jefferson’s DNA and compare it to available records, we only get a handful of matches that don’t tell a complete story. Let’s look at the first headline – was Jefferson Jewish? He didn’t practice Judaism and he wasn’t raised Jewish. He does have one genetic cousin who is a Moroccan Jew, but you have to go back about 2,000 years to find a common ancestor. While haplogroup T does have origins in the Middle-East, I wouldn’t say that it is definitely a Jewish haplogroup. If Jefferson were J1b2, there would be a stronger case tying him to the kohanim Jewish paternal lines. Jefferson also has a Belgian genetic cousin. Perhaps the headline should have been – Was Thomas Jefferson the first Belgian President? Not that exciting. Probably wouldn’t have sold very well.
Thomas Jefferson was a Phoenician! There are many articles attributing this statement to Spencer Wells as part of his In Search of Adam program in 2005. I can’t find one quote that actual has Wells saying this. In fact in 2008 Wells argued that the Phoenicians were haplogroup J2. Jefferson’s haplogroup T is found in the same places and at the same times as the Phoenician Mediterranean colonies. This may indicate that Jefferson’s ancestors travelled with the Phoenicians as a peer or as a slave. I don’t think that ethnicity by association works.
If Jefferson was a Phoenician, then so was Charlemagne. This is just plain and simple poor logic and a misunderstanding of genetics. As I mentioned, it doesn’t appear that haplogroup T is Phoenician. While Jefferson may be a descendant of Charlemagne, he is not a direct male descendant. You really need to be a direct male descendant to prove that an ancestor has the same y-DNA. One sample would never be enough to prove Charlemagne’s DNA. Multiple descendant samples and very strong genealogies are required to come close to determining an ancestor’s DNA. You never know where a non-paternal event may pop up.
Could Jefferson have recent origins in the Middle East? This writer never actually defines recent. We are left to wonder if the Jeffersons lied on their Naturalization applications. Based on ‘time to most recent common ancestor’ calculations, I’d put Jefferson’s ancestors in the Middle East about 3,000 years ago. I guess that’s fairly recent compared to the age of the universe.
Jefferson’s DNA traced to Egypt! One record match does not make an origin. That one Egyptian genetic cousin actually clusters better with other Moroccan records. This could indicate a back migration from Morocco to Egypt for that one person. A rule of thumb when determining origins is to find clusters of records. Jefferson does have a cluster on either side of the Strait of Gibraltar. This provides a strong argument that Jefferson’s ancestors came through that region and perhaps loitered there for a while. That doesn’t make it his origin.
How should we define Jefferson’s origins? It is important to define origins with context. Where in Britain did the Jeffersons come from? One biographer puts Jefferson’s family origins in Wales. Jefferson’s closest British genetic cousin comes from Yorkshire and the Jefferson surname has the highest distribution in Yorkshire. We are still talking about a single point of reference, so I won’t fall into the same trap and pronounce Thomas Jefferson a Yorkie. I can say that it appears that the Jeffersons were British and that the family had been in Britain for at least a 1,000 years. There’s just not enough data to be more certain.
Jefferson’s tribal DNA does leave a sparse trail of breadcrumbs across Europe in the 1,500 to 2,000 years ago range. There are genetic matches that become increasingly more distant in Belgium, France and Spain. We could connect-the-dots and we probably wouldn’t be far off the migration path. For Jefferson’s European origins, we might say his ancestors were Iberian. If we go back another 500 years, the picture changes to a culture that traveled the Mediterranean. The genetic breadcrumbs are in Morocco, Sicily, Cyprus, Egypt and Turkey. We could talk about Jefferson’s haplogroup T origins. A cluster of data suggests a southern Arabian Peninsula origin about 8,000 years ago.
If we continue backward in time, Jefferson’s ancestors came from East Africa just like everyone else on the planet. Which ‘origin’ you choose for Jefferson is completely up to your specific agenda. I use genetic genealogy to get a better understanding of the world that we live in and the things we have in common as one species. We may learn through DNA that our ancestors sacked Rome or pillaged the coast of England. That’s history, that’s fascinating, but that’s not who we are today. Unless you personally choose to embrace that history. Jefferson’s distant ancestors may have been Jewish, Phoenician or Egyptian, but that’s not who he was.
We shouldn’t persecute for the sins of our ancestors or sit on the laurels of their accomplishments. We need to keep moving forward in a positive direction.
#gDNA
Thomas Jefferson was a Phoenician! There are many articles attributing this statement to Spencer Wells as part of his In Search of Adam program in 2005. I can’t find one quote that actual has Wells saying this. In fact in 2008 Wells argued that the Phoenicians were haplogroup J2. Jefferson’s haplogroup T is found in the same places and at the same times as the Phoenician Mediterranean colonies. This may indicate that Jefferson’s ancestors travelled with the Phoenicians as a peer or as a slave. I don’t think that ethnicity by association works.
If Jefferson was a Phoenician, then so was Charlemagne. This is just plain and simple poor logic and a misunderstanding of genetics. As I mentioned, it doesn’t appear that haplogroup T is Phoenician. While Jefferson may be a descendant of Charlemagne, he is not a direct male descendant. You really need to be a direct male descendant to prove that an ancestor has the same y-DNA. One sample would never be enough to prove Charlemagne’s DNA. Multiple descendant samples and very strong genealogies are required to come close to determining an ancestor’s DNA. You never know where a non-paternal event may pop up.
Could Jefferson have recent origins in the Middle East? This writer never actually defines recent. We are left to wonder if the Jeffersons lied on their Naturalization applications. Based on ‘time to most recent common ancestor’ calculations, I’d put Jefferson’s ancestors in the Middle East about 3,000 years ago. I guess that’s fairly recent compared to the age of the universe.
Jefferson’s DNA traced to Egypt! One record match does not make an origin. That one Egyptian genetic cousin actually clusters better with other Moroccan records. This could indicate a back migration from Morocco to Egypt for that one person. A rule of thumb when determining origins is to find clusters of records. Jefferson does have a cluster on either side of the Strait of Gibraltar. This provides a strong argument that Jefferson’s ancestors came through that region and perhaps loitered there for a while. That doesn’t make it his origin.
How should we define Jefferson’s origins? It is important to define origins with context. Where in Britain did the Jeffersons come from? One biographer puts Jefferson’s family origins in Wales. Jefferson’s closest British genetic cousin comes from Yorkshire and the Jefferson surname has the highest distribution in Yorkshire. We are still talking about a single point of reference, so I won’t fall into the same trap and pronounce Thomas Jefferson a Yorkie. I can say that it appears that the Jeffersons were British and that the family had been in Britain for at least a 1,000 years. There’s just not enough data to be more certain.
Jefferson’s tribal DNA does leave a sparse trail of breadcrumbs across Europe in the 1,500 to 2,000 years ago range. There are genetic matches that become increasingly more distant in Belgium, France and Spain. We could connect-the-dots and we probably wouldn’t be far off the migration path. For Jefferson’s European origins, we might say his ancestors were Iberian. If we go back another 500 years, the picture changes to a culture that traveled the Mediterranean. The genetic breadcrumbs are in Morocco, Sicily, Cyprus, Egypt and Turkey. We could talk about Jefferson’s haplogroup T origins. A cluster of data suggests a southern Arabian Peninsula origin about 8,000 years ago.
If we continue backward in time, Jefferson’s ancestors came from East Africa just like everyone else on the planet. Which ‘origin’ you choose for Jefferson is completely up to your specific agenda. I use genetic genealogy to get a better understanding of the world that we live in and the things we have in common as one species. We may learn through DNA that our ancestors sacked Rome or pillaged the coast of England. That’s history, that’s fascinating, but that’s not who we are today. Unless you personally choose to embrace that history. Jefferson’s distant ancestors may have been Jewish, Phoenician or Egyptian, but that’s not who he was.
We shouldn’t persecute for the sins of our ancestors or sit on the laurels of their accomplishments. We need to keep moving forward in a positive direction.
#gDNA
Subscribe to:
Posts (Atom)












.png)
.png)


